Sohl Group Page
Curriculum Vitae
- B.S., Biochemistry, summa cum laude, University of Oklahoma, 2005
- Ph.D., Biochemistry, Vanderbilt University, 2010
- NIH Postdoctoral Fellow, Yale University, 2010-2015
- Assistant Professor, San Diego State University 2015-2021
- Associate Professor, San Diego State University 2021-present
Research Interests
In the Sohl lab, we are interested in probing mechanistic questions at the intersection of biochemistry and human disease. We use kinetic, structural, and cellular tools to address how altered enzyme activity impacts human health. In particular, we are interested in exploring the catalytic, structural and functional consequences of enzyme mutations implicated in diseases such as cancer. By understanding these molecular mechanisms of dysfunction, we can illuminate structure-function relationships, identify affected downstream pathways and ultimately develop platforms for targeted therapy. Students in the Sohl lab will gain interdisciplinary expertise in biochemistry, molecular biology and biophysics, including pre-steady-state kinetics, X-ray crystallography, spectroscopy, recombinant technology and cellular methods in a collaborative environment. We are recruiting inquisitive, dedicated and passionate students and postdocs to help us make a meaningful impact on improving human health.
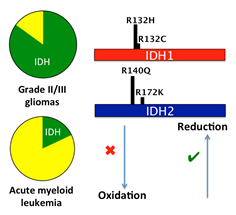
Enzymatic mechanisms of cancer. Enzyme dysfunction can result from mutation, misregulation or amplification, which can lead to cancer and other diseases. In particular, a mechanistic understanding of altered enzyme function provides a critical foundation for understanding tumorigenesis. One focus of the Sohl lab is elucidating the catalytic pathway and structural features of isocitrate dehydrogenase (IDH) mutations that have been implicated in gliomas and leukemia. Some IDH mutations are unique in that they have the potential to confer both oncogenic and tumor suppressive properties, and can even result in generation of a novel oncometabolite product. This hints that intriguing and complex catalytic and structural alterations must be at work. A second focus of the lab is exploring the role of kinases in diseases such as uterine and stomach cancers and venous malformations. Kinases are the most common class of enzymes to be implicated in cancer, often through activating mutations. By exploring the catalytic and structural consequences of these mutations and the resulting effects on downstream pathways, we can better understand oncogenesis. Ultimately, these studies can provide important tools for target identification and drug development.
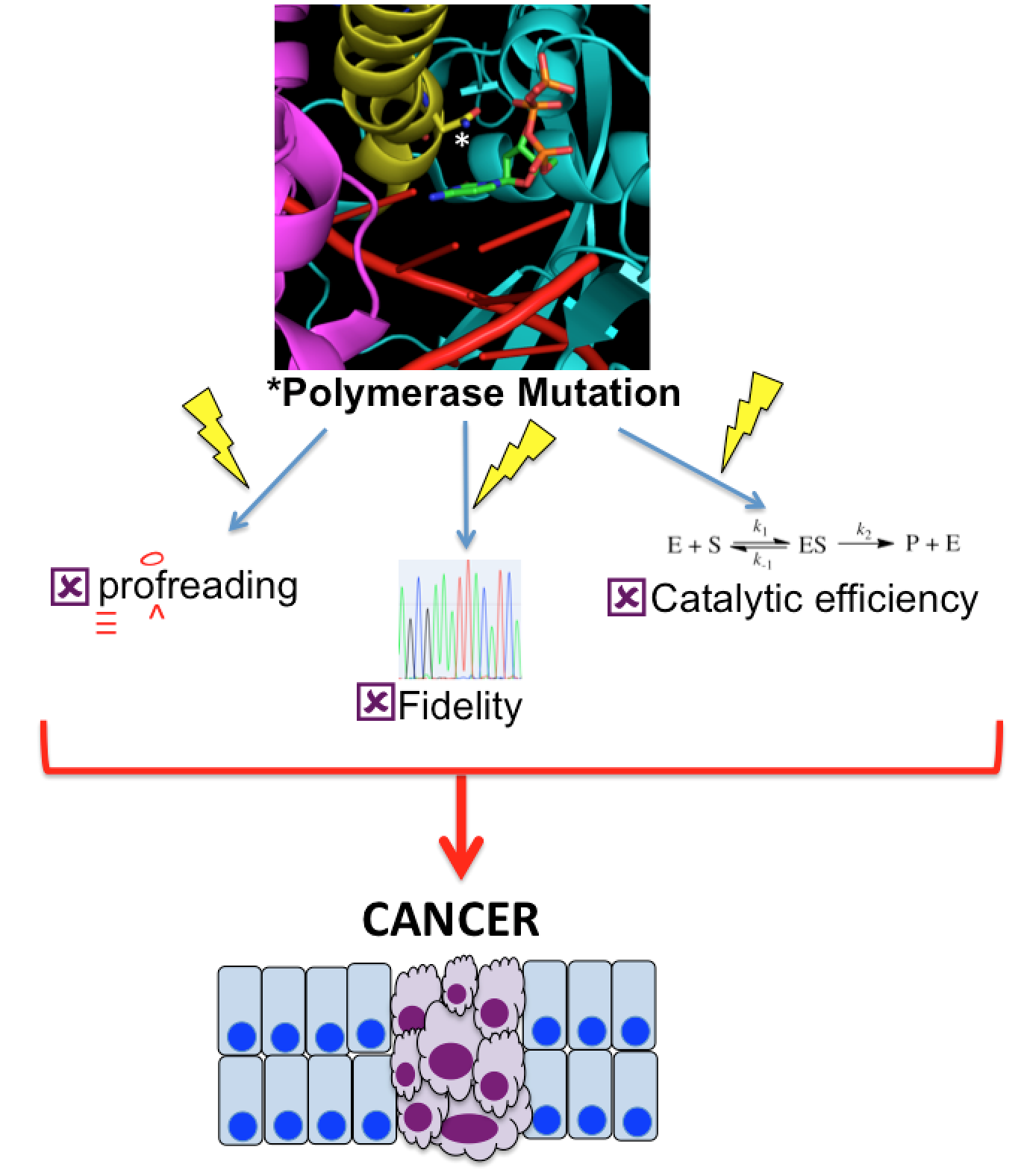
The many paths to genome instability. Mutations can alter the activity of human polymerases, which in turn can lead to cancer and rare congenital diseases. Our goal is to understand how these mutations alter the ability of polymerases to catalyze efficient genome replication. Importantly, proposed tumor suppressing mutations in A and B family DNA polymerases have the potential to lead to tumorigenesis through vastly different mechanisms, including changing rates of polymerization or altering fidelity, and these mechanisms in turn can use a variety of approaches to achieve these devastating outcomes. We are interested in exploring the molecular mechanisms of tumor suppressive polymerase mutations and identifying the downstream affects on DNA damage and repair. This will help provide tools for assessing DNA damage repair pathways as therapeutic targets.
Selected Publications
- Professor Sohl's full publications list at the NCBI.
- "Active Site Remodeling in Tumor-relevant {IDH1} Mutants Drives Distinct Kinetic Features and Potential Resistance Mechanisms,"
M. Mealka, N. A. Sierra, D. Avellaneda Matteo, E. Albekioni, R. Khoury, T. Mai, B. M. Conley, N. J. Coleman, K. A. Sabo, E. A. Komives, A. A. Bobkov, A. L. Cooksy, S. Silletti, J. M. Schiffer, T. Huxford, C. D. Sohl,
Nature Commun. 15, 3785 (2024). (doi: 10.1038/s41467-024-48277-2.) - "Capturing the Dynamic Conformational Changes of Human Isocitrate Dehydrogenase 1 (IDH1) upon Ligand and Metal Binding Using Hydrogen-Deuterium Exchange Mass Spectrometry,"
Sabo K.A., Albekioni E., Caliger D., Coleman N.J., Thornberg E., Avellaneda Matteo D., Komives E.A., Silletti S., Sohl C.D.,
Biochemistry 62, 1145-1159 (2023). (doi: 10.1021/acs.biochem.2c00636.) - "Insights into Nitrosoalkane Binding to Myoglobin Provided by Crystallography of Wild-Type and Distal Pocket Mutant Derivatives,"
Herrera V.E., Charles T.P., Scott T.G., Prather K.Y., Nguyen N.T., Sohl C.D., Thomas L.M., Richter-Addo G.B.,
Biochemistry 62, 1406-1419 (2023). (doi: 10.1021/acs.biochem.2c00725.) - "Probing altered enzyme activity in the biochemical characterization of cancer,"
Adam M.A.A., Sohl C.D.,
Bioscience Reports 42 (2), BSR20212002 (2022). (doi: 10.1042/BSR20212002.) - "Readying students for careers in industry: A guided inquiry activity to prepare students for success in biotechnology and pharmaceutical industry positions,"
Gross S., Sohl C.D.,
Biochemistry and Molecular Biology Education 49, 407-415 (2021). (doi: 10.1002/bmb.21491.) - "Evaluating mechanisms of idh1 regulation through site-specific acetylation mimics,"
Weeks J., Strom A.I., Widjaja V., Alexander S., Pucher D.K., Sohl C.D.,
Biomolecules 11, 740 (2021). (doi: 10.3390/biom11050740.) - "An acidic residue buried in the dimer interface of isocitrate dehydrogenase 1 (IDH1) helps regulate catalysis and pH sensitivity,"
Luna L.A., Lesecq Z., White K.A., Hoang A., Scott D.A., Zagnitko O., Bobkov A.A., Barber D.L., Schiffer J.M., Isom D.G., Sohl C.D.,
Biochemical Journal 477, 2999-3018 (2020). (doi: 10.1042/BCJ20200311.) - "Water Networks and Correlated Motions in Mutant Isocitrate Dehydrogenase 1 (IDH1) Are Critical for Allosteric Inhibitor Binding and Activity,"
Jennifer M. Chambers, Wade Miller, Giovanni Quichocho, Viraj Upadhye, Diego Avellaneda Matteo, Andrey A. Bobkov, Christal D. Sohl, and Jamie M. Schiffer,
Biochem. 59, 479-490 (2020). (10.1021/acs.biochem.9b01023.) - "Activity of Selected Nucleoside Analogue ProTides against Zika Virus in Human Neural Stem Cells,"
Jean A. Bernatchez, Michael Coste, Sungjun Beck, Grace A. Wells, Lucas A. Luna, Alex E. Clark, Zhe Zhu, David Hecht, Jeremy N. Rich, Christal D. Sohl, Byron W. Purse, and Jair L. Siqueira-Neto,
Viruses 11, 365 (2019). (doi: 10.3390/v11040365.) - "A Tie2 kinase mutation causing venous malformations increases phosphorylation rates and enhances cooperativity,"
Madison A. Kennedy, Zeqing Xu, Yunjin Wu, and Christal D.Sohl,
Biochem. Biophys. Res. Comm. 509, 898-902 (2019). (doi: 10.1016/j.bbrc.2019.01.020.) - "The FGFR1 V561M Gatekeeper Mutation Drives AZD4547 Resistance through STAT3 Activation and EMT,"
Molly R. Ryan, Christal D. Sohl, BeiBei Luo, and Karen S. Anderson,
Molec. Cancer Res. 17, 532-543 (2019). (doi: 10.1158/1541-7786.MCR-18-0429.)